Bio-theory Lab Notes!
Growth Rates and Worm Posture
An update on my explorations this weekend, applying models to biological data. Here are some rough lab notes I’ve made.
Bioreactor fermentation
Thanks to David Jordan at Living Physics for giving me some of his growth datasets from the lab’s bioreactors!
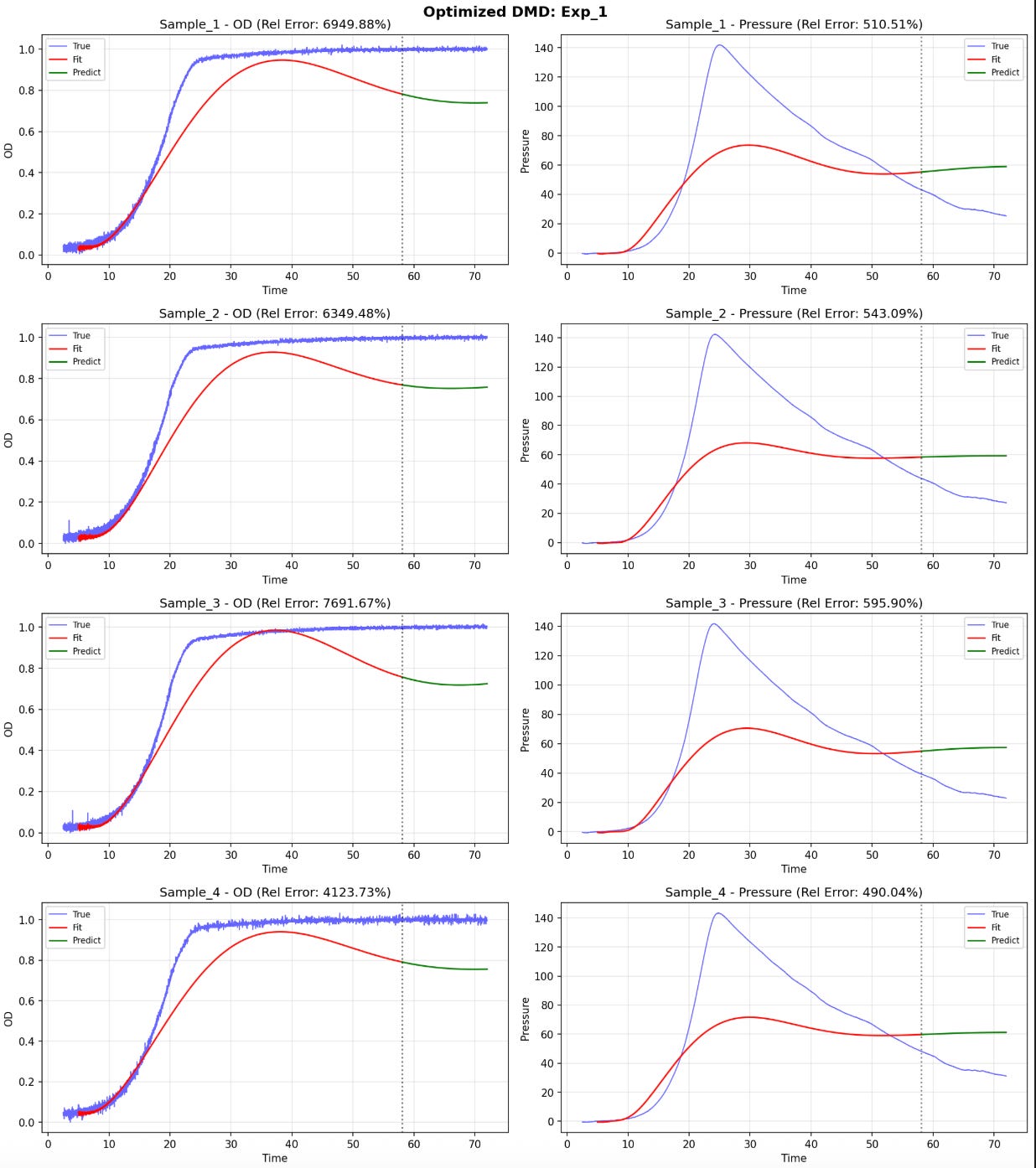
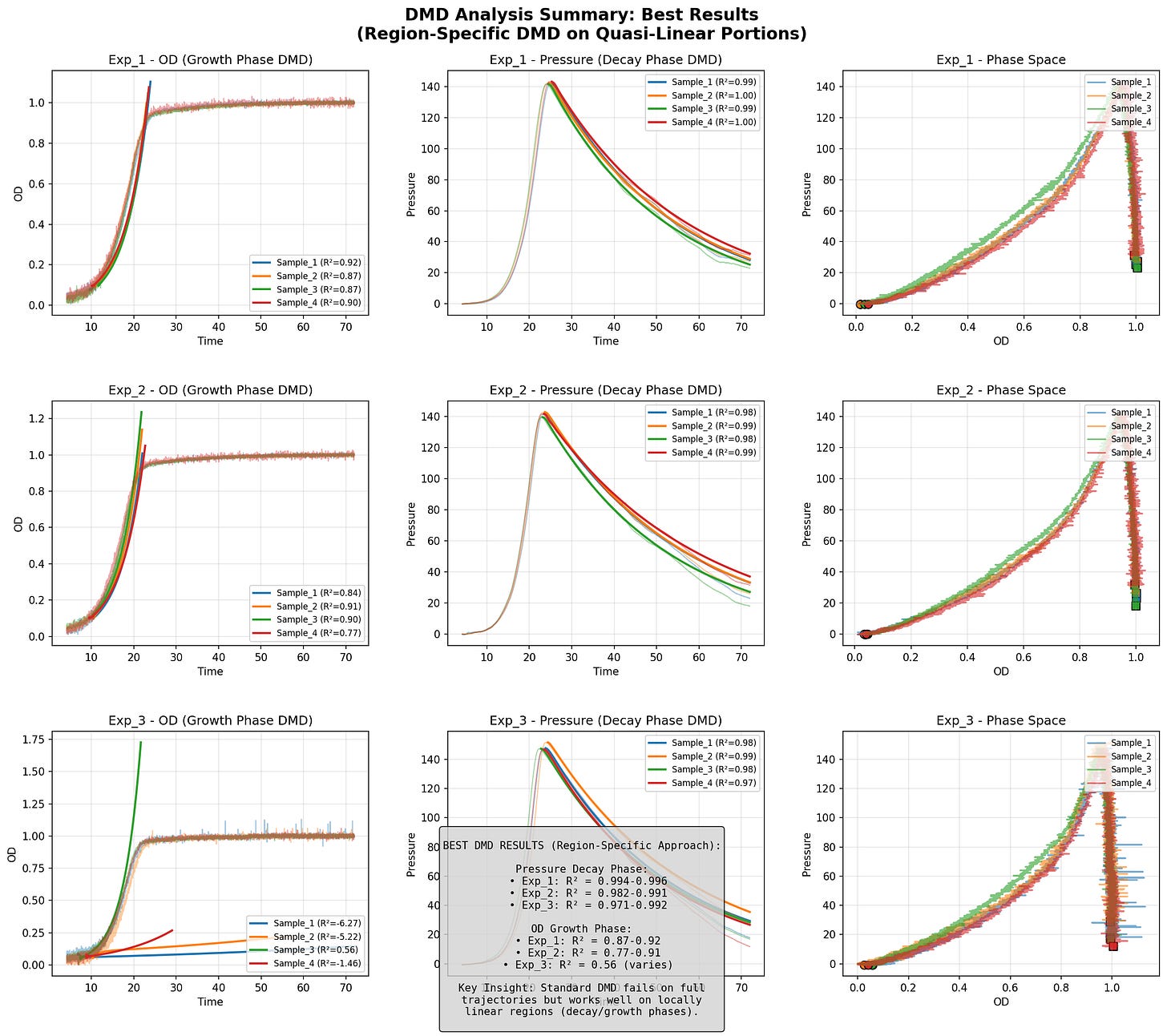
The data consisted of 3 experiments, 4 samples each. The two variables measured were OD (cell growth) and pressure over time.
I tried to apply Hankel-DMD with 300 delays, rank 80, to model the change in these variables over time.
I struggled to get models to capture the full dynamics. But it works okay on the two regimes separately. DMD captures growth phase well. Pressure decay phase hits R² ~0.88-0.99. Standard DMD fails on full trajectories but works on quasi-linear portions.
Worm Posture Data
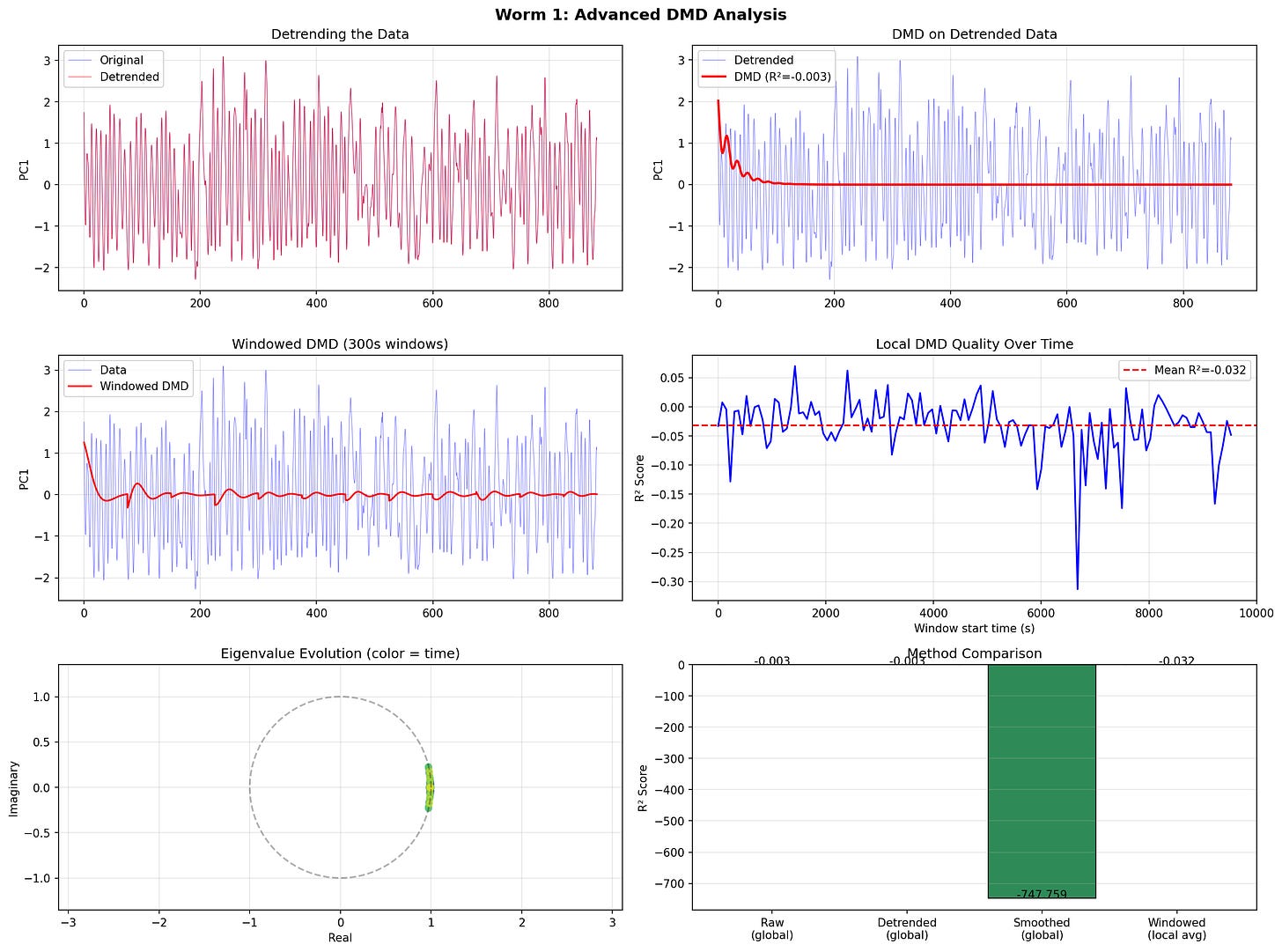
David also got me some data on C. Elegans (roundworm) reduced posture data.
I threw four DMD variants at them: standard, Hankel, extended, windowed, which didn’t work so well.
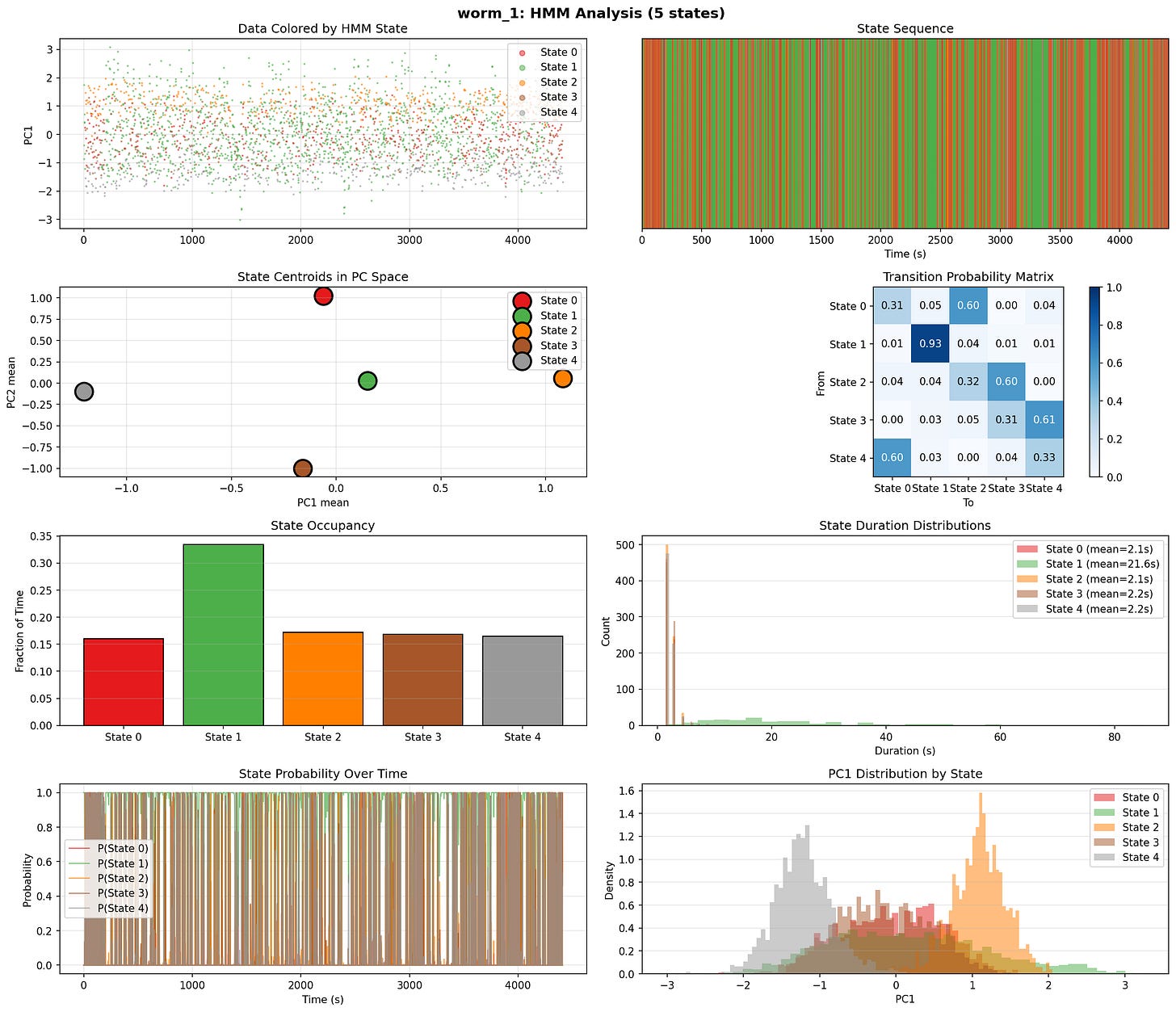
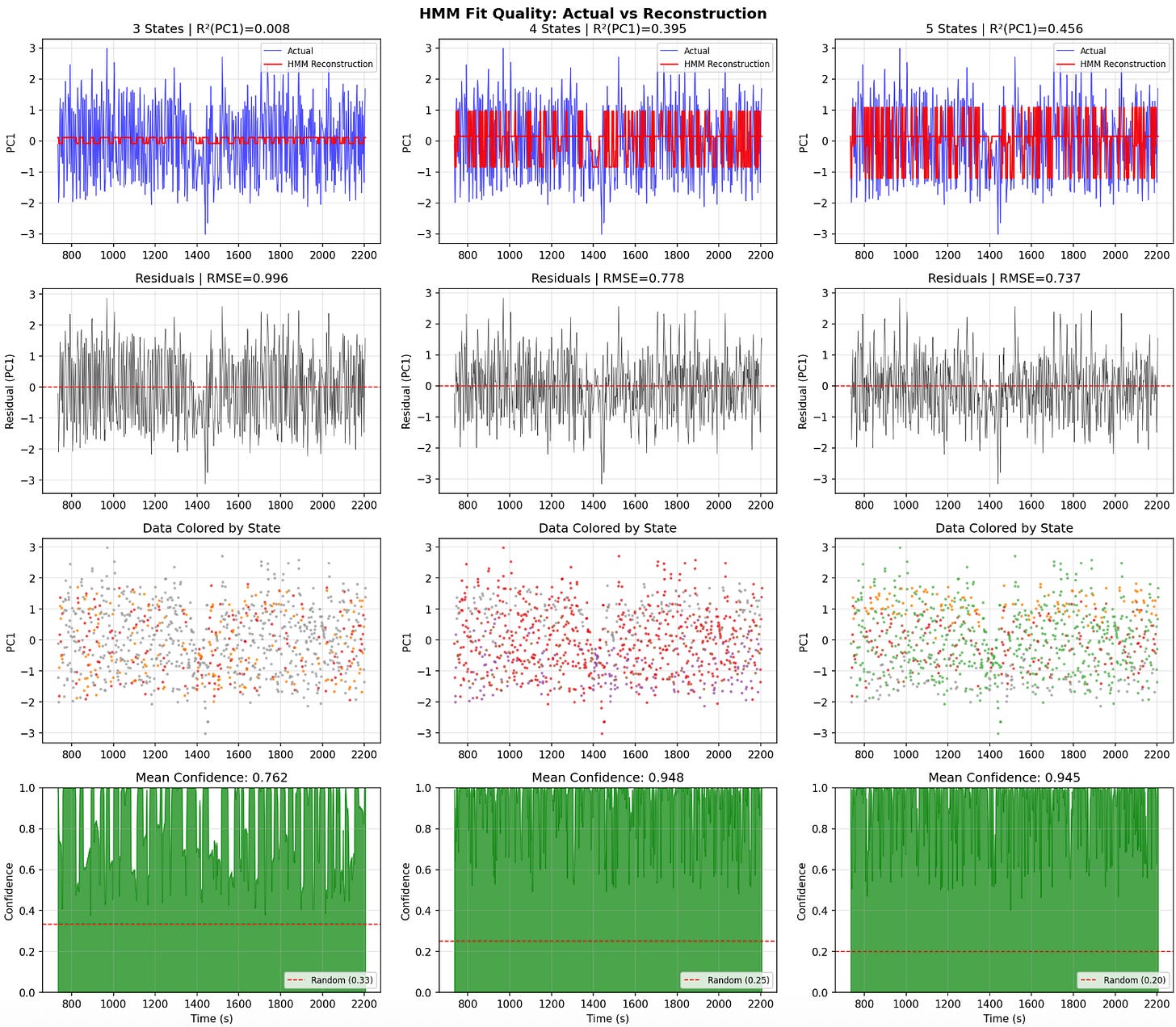
So I ran ran hidden markov moels to try and find discrete behavioral states
5-state HMM gives best fit - and it worked quite well - need to dig into this more.
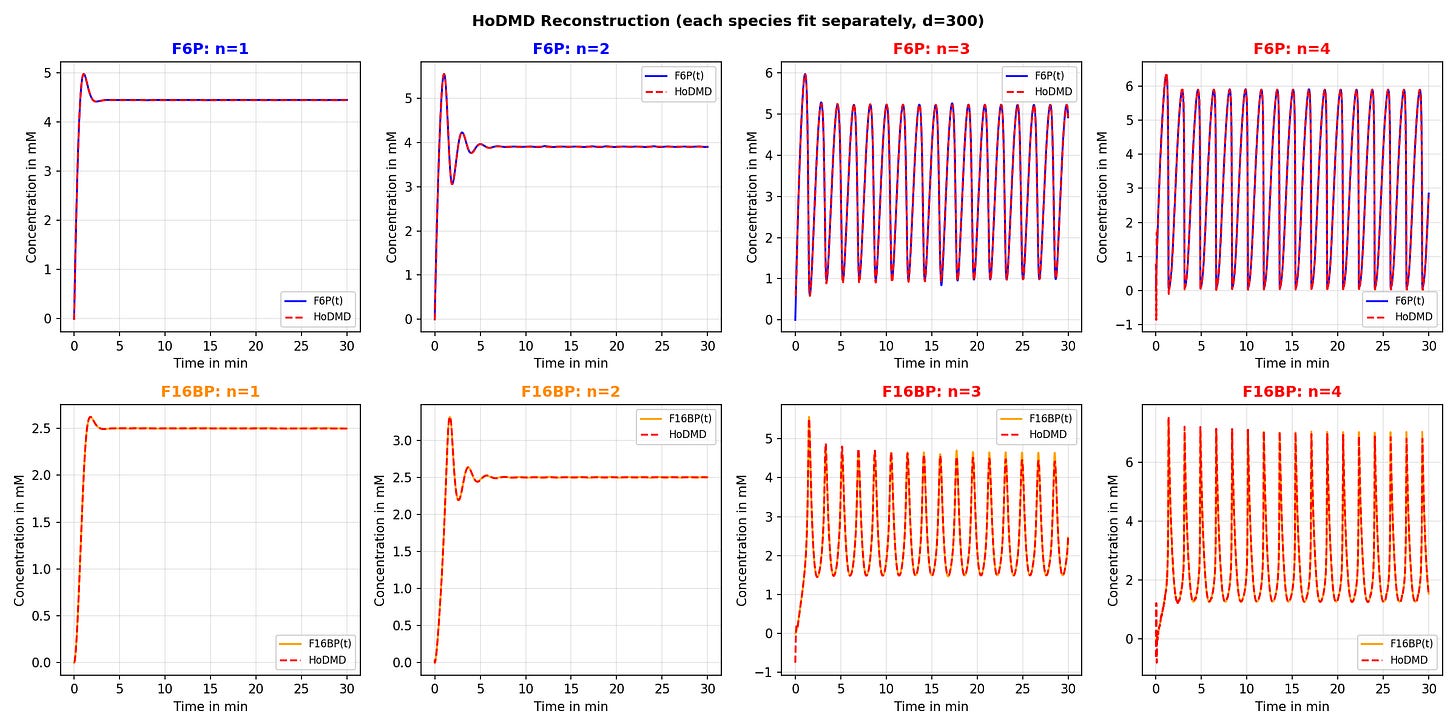
Glycolysis oscillations
Tried replicating glycolysis reaction simulations in Wüstner et al. (2025) — they claim HoDMD with d=300 gives “exact” reconstruction - seems to check out, pretty keen on exploring this further.